einstein (São Paulo). 01/Jul/2014;12(3):336-41.
Transcriptional expression study in the central nervous system of rats: what gene should be used as internal control?
DOI: 10.1590/S1679-45082014AO3042
Objective
A growing number of published articles report the expression of specific genes with different behavior patterns in rats. The levels of messenger ribonucleic acid transcripts are usually analyzed by reverse transcription followed by polymerase chain reaction and quantified after normalization with an internal control or reference gene (housekeeping gene). Nevertheless, housekeeping genes exhibit different expression in the central nervous system, depending on the physiological conditions and the area of the brain to be studied. The choice of a good internal control gene is essential for obtaining reliable results. This study evaluated the expression of three housekeeping genes (beta-actin, cyclophilin A, and ubiquitin C) in different areas of the central nervous system in rats (olfactory bulb, hippocampus, striatum, and prefrontal cortex).
Methods
Wistar rats (virgin females, n=6) during the diestrum period were used. Total ribonucleic acid was extracted from each region of the brain; the complementary deoxyribonucleic acid was synthesized by reverse transcription and amplified by real-time quantitative polymerase chain reaction using SYBR™ Green and primers specific for each one of the reference genes. The stability of the expression was determined using NormFinder.
Results
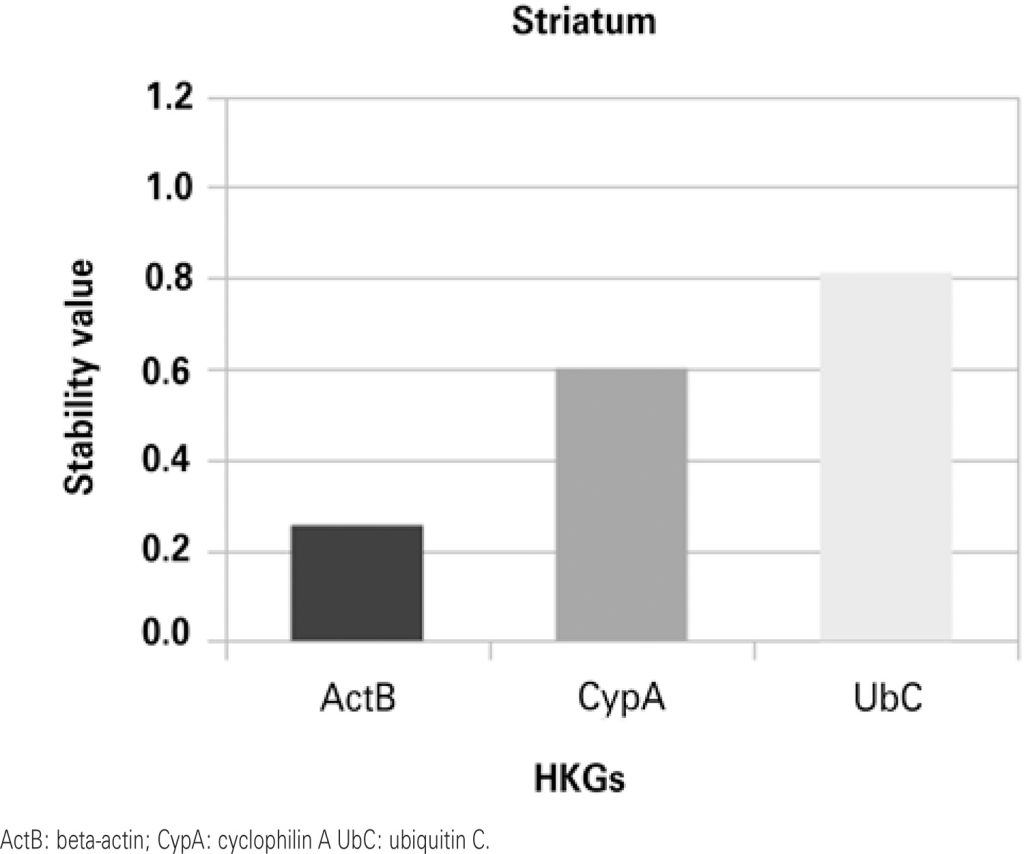
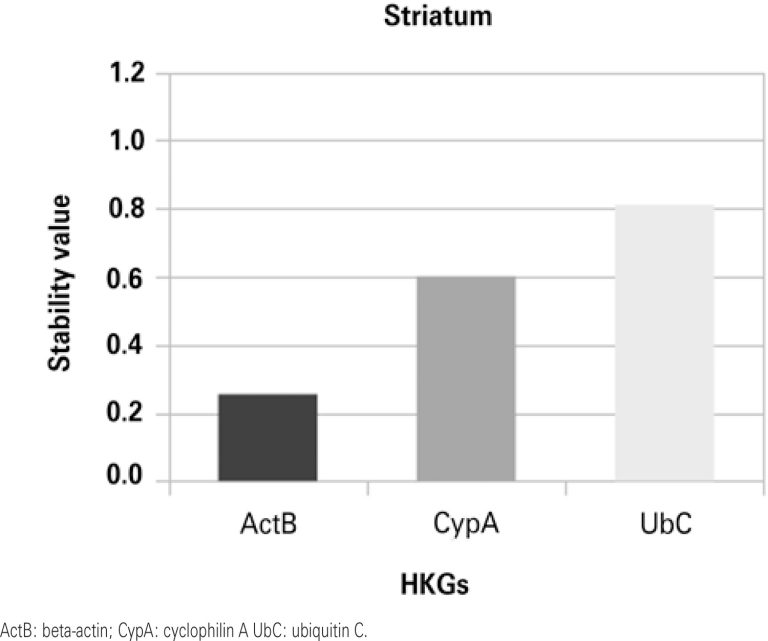
Beta-actin was the most stable gene in the hippocampus and striatum, while cyclophilin A and ubiquitin C showed greater stability in the prefrontal cortex and the olfactory bulb, respectively.
Conclusion
Based on our study, further studies of gene expression using rats as animal models should take into consideration these results when choosing a reliable internal control gene.
153